2IVS - chain B (model B) | Ret proto-oncogene
Structure information
| PDB: | 2IVS |
| PubMed: | 16928683 |
| Release date: | 2006-08-14 |
| Resolution: | 2 Å |
| Kinase: | RET |
| Family: | Ret |
| Group: | TK |
| Species: | HUMAN |
| Quality Score: | 8.3 |
| Missing Residues: | 0 |
| Missing Atoms: | 17 |
| DFG conformation: | in |
| αC-helix conformation: | in |
| Salt bridge KIII.17 and EαC.24: | Yes (2.9Å) |
| ASP rotation (xDFG.81) : | 340° |
| PHE rotation (xDFG.82) : | 20° |
| Activation loop position: | -3.2Å |
| αC-helix position: | 16.5Å |
| G-rich loop angle: | 52.2° |
| G-rich loop distance: | 15.2Å |
| G-rich loop rotation: | 67.9° |
Other models from this PDB:
2D & 3D views
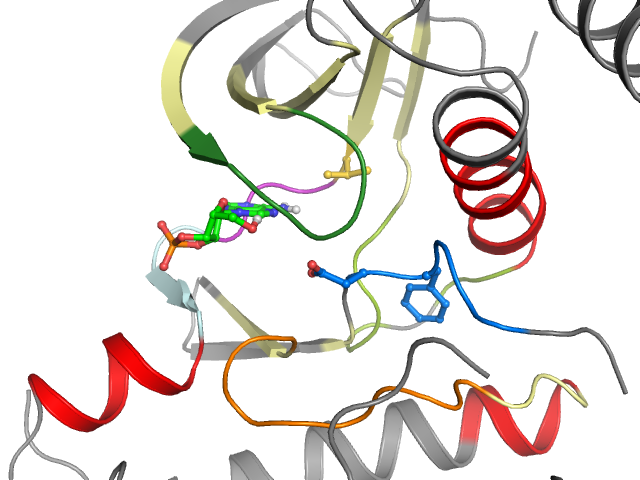
Binding pocket waters
The following waters were found in the defined clusters:
I5
H-bond protein
I6
H-bond ligand
H-bond protein
I7
H-bond ligand
Binding pocket sequence
| Uniprot | KTLGEGEFGKVVKVAVKMLDLLSEFNVLKQVNPHVIKLYGALLIVEYAKYGSLRGFLREYLAEMKLVHRDLAARNILVISDFGLS |
| Structure: | KTLGEGEFGKVVKVAVKMLDLLSEFNVLKQVNPHVIKLYGALLIVEYAKYGSLRGFLREYLAEMKLVHRDLAARNILVISDFGLS |
Modified residues
No modified residues identified.
Orthosteric ligand
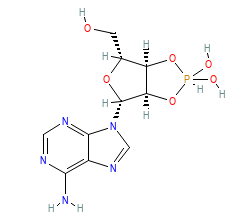
Ligand HET-code: ACK
Ligand Name: (3AR,4R,6R,6AR)-4-(6-AMINO-9H-PURIN-9-YL)-6-(HYDROXYMETHYL)TETRAHYDROFURO[3,4-D][1,3,2]DIOXAPHOSPHOL-2-OL 2-OXIDE
- Download image
- LABELS
- KLIFS residue #
- Amino Acid
- None
- COLORS
- Interaction types
- KLIFS (all res.)
- KLIFS (interacting res.)
- None
- OTHER
- Show/hide non-interacting res.
- En/disable resizing interacting res.
This ligand targets the following (sub)pockets:
| Main pockets | |
|---|---|
| Front | |
| Gate | |
| Back | |
| Subpockets | |
|---|---|
| FP-I | |
| FP-II | |
| BP-I-A | |
| BP-I-B | |
| BP-II-in | |
| BP-II-A-in | |
| BP-II-B-in | |
| BP-II-out | |
| BP-II-B | |
| BP-III | |
| BP-IV | |
| BP-V | |
Kinase-ligand interactions
■ Hydrophobic ♦ Aromatic face-to-face ♦ Aromatic face-to-edge ▲ H-bond donor ▲ H-bond acceptor ● Ionic positive ● Ionic negative
| I | g.l | II | III | αC | |||||||||||||||
| 1 K 728 | 2 T 729 | 3 L 730 | 4 G 731 | 5 E 732 | 6 G 733 | 7 E 734 | 8 F 735 | 9 G 736 | 10 K 737 | 11 V 738 | 12 V 739 | 13 K 740 | 14 V 755 | 15 A 756 | 16 V 757 | 17 K 758 | 18 M 759 | 19 L 760 | 20 D 771 |
| ■ | ■ | ■ | ■ | ■ | ■ | ||||||||||||||
| αC | b.l | IV | |||||||||||||||||
| 21 L 772 | 22 L 773 | 23 S 774 | 24 E 775 | 25 F 776 | 26 N 777 | 27 V 778 | 28 L 779 | 29 K 780 | 30 Q 781 | 31 V 782 | 32 N 783 | 33 P 785 | 34 H 786 | 35 V 787 | 36 I 788 | 37 K 789 | 38 L 790 | 39 Y 791 | 40 G 792 |
| IV | V | GK | hinge | linker | αD | αE | |||||||||||||
| 41 A 793 | 42 L 801 | 43 L 802 | 44 I 803 | 45 V 804 | 46 E 805 | 47 Y 806 | 48 A 807 | 49 K 808 | 50 Y 809 | 51 G 810 | 52 S 811 | 53 L 812 | 54 R 813 | 55 G 814 | 56 F 815 | 57 L 816 | 58 R 817 | 59 E 818 | 60 Y 864 |
| ▲ | ■♦ | ■▲ | ▲ | ||||||||||||||||
| αE | VI | c.l | VII | VIII | x | ||||||||||||||
| 61 L 865 | 62 A 866 | 63 E 867 | 64 M 868 | 65 K 869 | 66 L 870 | 67 V 871 | 68 H 872 | 69 R 873 | 70 D 874 | 71 L 875 | 72 A 876 | 73 A 877 | 74 R 878 | 75 N 879 | 76 I 880 | 77 L 881 | 78 V 882 | 79 I 890 | 80 S 891 |
| ■ | |||||||||||||||||||
| DFG | a.l | ||||||||||||||||||
| 81 D 892 | 82 F 893 | 83 G 894 | 84 L 895 | 85 S 896 | |||||||||||||||
Binding affinities
Ligand not found in ChEMBL.