1YOL - chain A | SRC proto-oncogene, non-receptor tyrosine kinase
Structure information
| PDB: | 1YOL |
| PubMed: | 16168436 |
| Release date: | 2006-01-27 |
| Resolution: | 2.3 Å |
| Kinase: | SRC |
| Family: | Src |
| Group: | TK |
| Species: | HUMAN |
| Quality Score: | 6.5 |
| Missing Residues: | 6 |
| Missing Atoms: | 7 |
| DFG conformation: | na |
| αC-helix conformation: | in |
| Salt bridge KIII.17 and EαC.24: | No |
| ASP rotation (xDFG.81) : | ° |
| PHE rotation (xDFG.82) : | 180° |
| Activation loop position: | 0Å |
| αC-helix position: | 16.3Å |
| G-rich loop angle: | 58.1° |
| G-rich loop distance: | 18.2Å |
| G-rich loop rotation: | 52.7° |
Other models from this PDB:
2D & 3D views
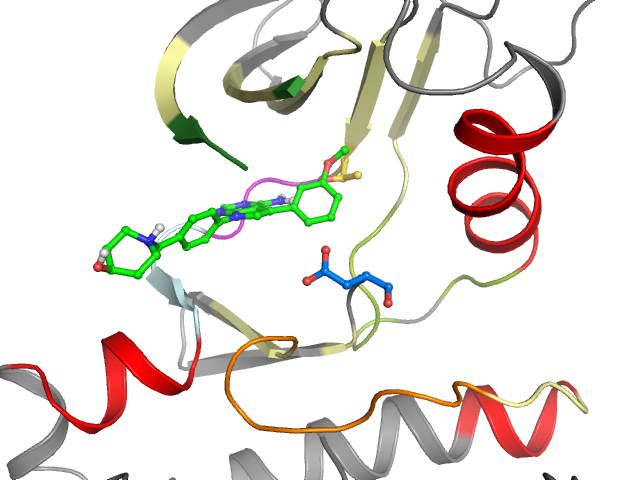
Binding pocket waters
The following waters were found in the defined clusters:
I1
H-bond protein
I2
H-bond protein
I7
H-bond ligand
Binding pocket sequence
| Uniprot | VKLGQGCFGEVWMVAIKTLAFLQEAQVMKKLREKLVQLYAVYIVTEYMSKGSLLDFLKGYVERMNYVHRDLRAANILVVADFGLA |
| Structure: | VKLGQG__GEVWMVAIKTLAFLQEAQVMKKLREKLVQLYAVYIVTEYMNKGSLLDFLKGYVERMNYVHRDLRAANILVVAD____ |
Modified residues
No modified residues identified.
Orthosteric ligand
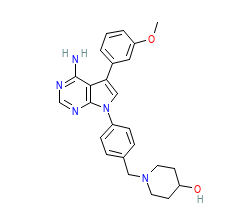
Ligand HET-code: S03
Ligand Name: 1-{4-[4-AMINO-5-(3-METHOXYPHENYL)-7H-PYRROLO[2,3-D]PYRIMIDIN-7-YL]BENZYL}PIPERIDIN-4-OL
- Download image
- LABELS
- KLIFS residue #
- Amino Acid
- None
- COLORS
- Interaction types
- KLIFS (all res.)
- KLIFS (interacting res.)
- None
- OTHER
- Show/hide non-interacting res.
- En/disable resizing interacting res.
This ligand targets the following (sub)pockets:
| Main pockets | |
|---|---|
| Front | |
| Gate | |
| Back | |
| Subpockets | |
|---|---|
| FP-I | |
| FP-II | |
| BP-I-A | |
| BP-I-B | |
| BP-II-in | |
| BP-II-A-in | |
| BP-II-B-in | |
| BP-II-out | |
| BP-II-B | |
| BP-III | |
| BP-IV | |
| BP-V | |
Kinase-ligand interactions
■ Hydrophobic ♦ Aromatic face-to-face ♦ Aromatic face-to-edge ▲ H-bond donor ▲ H-bond acceptor ● Ionic positive ● Ionic negative
| I | g.l | II | III | αC | |||||||||||||||
| 1 V 273 | 2 K 274 | 3 L 275 | 4 G 276 | 5 Q 277 | 6 G 278 | 7 _ _ | 8 _ _ | 9 G 281 | 10 E 282 | 11 V 283 | 12 W 284 | 13 M 285 | 14 V 294 | 15 A 295 | 16 I 296 | 17 K 297 | 18 T 298 | 19 L 299 | 20 A 308 |
| ■ | ■ | ■ | ■ | ■ | ■ | ||||||||||||||
| αC | b.l | IV | |||||||||||||||||
| 21 F 309 | 22 L 310 | 23 Q 311 | 24 E 312 | 25 A 313 | 26 Q 314 | 27 V 315 | 28 M 316 | 29 K 317 | 30 K 318 | 31 L 319 | 32 R 320 | 33 E 322 | 34 K 323 | 35 L 324 | 36 V 325 | 37 Q 326 | 38 L 327 | 39 Y 328 | 40 A 329 |
| IV | V | GK | hinge | linker | αD | αE | |||||||||||||
| 41 V 330 | 42 Y 337 | 43 I 338 | 44 V 339 | 45 T 340 | 46 E 341 | 47 Y 342 | 48 M 343 | 49 N 344 | 50 K 345 | 51 G 346 | 52 S 347 | 53 L 348 | 54 L 349 | 55 D 350 | 56 F 351 | 57 L 352 | 58 K 353 | 59 G 354 | 60 Y 378 |
| ■ | ■ | ■ | ▲ | ■♦ | ■▲ | ■ | ■ | ■ | |||||||||||
| αE | VI | c.l | VII | VIII | x | ||||||||||||||
| 61 V 379 | 62 E 380 | 63 R 381 | 64 M 382 | 65 N 383 | 66 Y 384 | 67 V 385 | 68 H 386 | 69 R 387 | 70 D 388 | 71 L 389 | 72 R 390 | 73 A 391 | 74 A 392 | 75 N 393 | 76 I 394 | 77 L 395 | 78 V 396 | 79 V 404 | 80 A 405 |
| ■ | ■ | ||||||||||||||||||
| DFG | a.l | ||||||||||||||||||
| 81 D 406 | 82 _ _ | 83 _ _ | 84 _ _ | 85 _ _ | |||||||||||||||
| ■ | |||||||||||||||||||
Binding affinities
Ligand not found in ChEMBL.