4EBV - chain A (model B) | Protein tyrosine kinase 2
Structure information
| PDB: | 4EBV |
| PubMed: | 22819505 |
| Release date: | 2012-08-22 |
| Resolution: | 1.67 Å |
| Kinase: | PTK2 (FAK) |
| Family: | FAK |
| Group: | TK |
| Species: | HUMAN |
| Quality Score: | 5.6 |
| Missing Residues: | 3 |
| Missing Atoms: | 4 |
| DFG conformation: | out-like |
| αC-helix conformation: | in |
| Salt bridge KIII.17 and EαC.24: | Yes (2.8Å) |
| ASP rotation (xDFG.81) : | 125° |
| PHE rotation (xDFG.82) : | 236° |
| Activation loop position: | 3.1Å |
| αC-helix position: | 18Å |
| G-rich loop angle: | 39.1° |
| G-rich loop distance: | 13.2Å |
| G-rich loop rotation: | 129.5° |
Other models from this PDB:
2D & 3D views
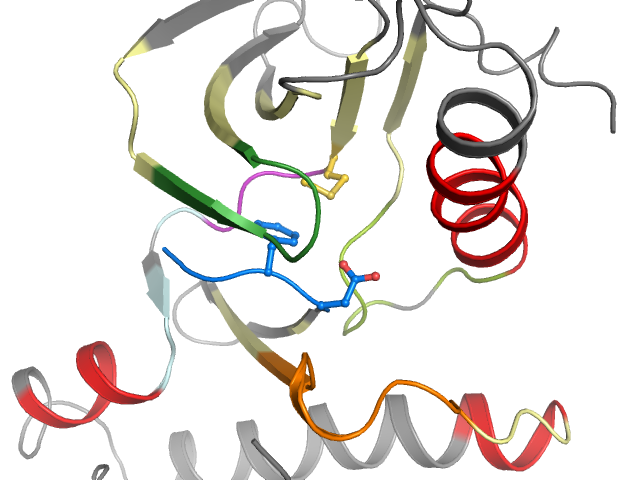
Binding pocket waters
The following waters were found in the defined clusters:
I4
H-bond protein
I8
H-bond protein
Binding pocket sequence
| Uniprot | RCIGEGQFGDVHQVAIKTCKFLQEALTMRQFDPHIVKLIGVWIIMELCTLGELRSFLQVYLESKRFVHRDIAARNVLVLGDFGLS |
| Structure: | RCIGEGQFGDVHQVAIKTCKFLQEALTMRQFDPHIVKLIGVWIIMELCTGELR__SFLQYLESKRFVHRDIAARNVLVLGDFGL_ |
Modified residues
No modified residues identified.
Allosteric ligand
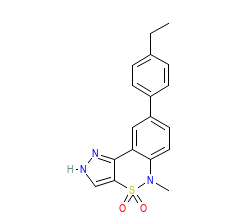
Ligand HET-code: 0O7
Ligand Name: 8-(4-ethylphenyl)-5-methyl-2,5-dihydropyrazolo[4,3-c][2,1]benzothiazine 4,4-dioxide
Binding affinities
Ligand not found in ChEMBL.